Home / Introduction to Peptide Synthesis
Amines
Introduction to Peptide Synthesis
Last updated: August 27th, 2025 |
Peptide bonds: Forming peptides from amino acids with the use of protecting groups
Today we’ll go deeper on how to synthesize the most important amides of all – peptides – with an important contribution from protecting group chemistry.
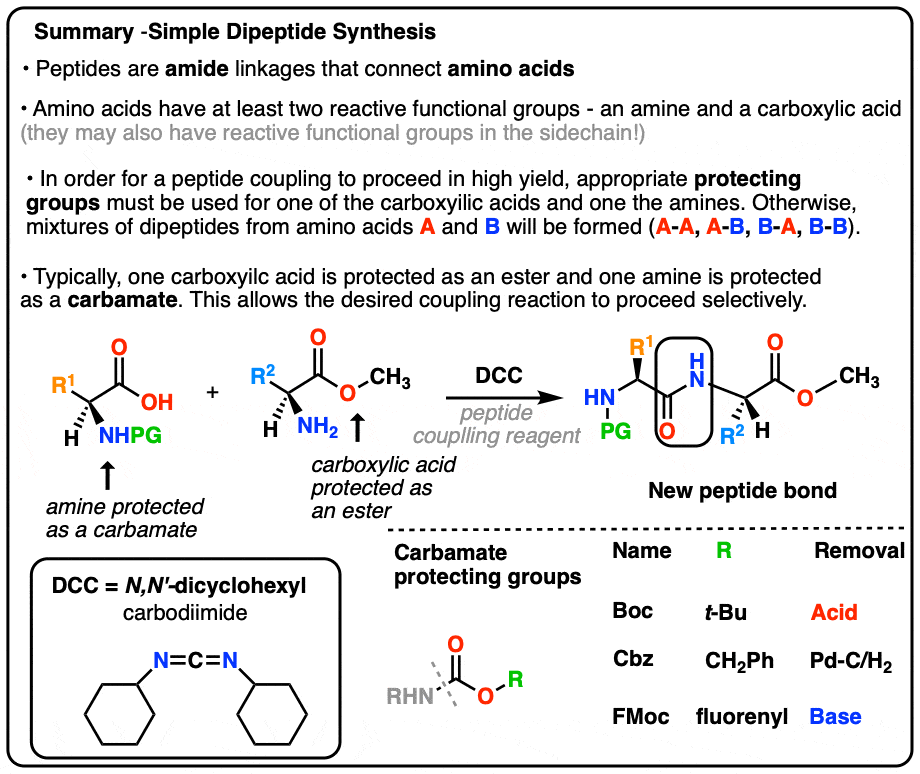
Table of Contents
- What Are Peptide Bonds?
- The “Proteinogenic” Amino Acids
- Synthesis of a Dipeptide Without Protecting Groups
- Synthesis of a Dipeptide Using A Protecting Group Strategy
- Synthesis of Longer Peptides – Tripeptides and Tetrapeptides
- Bonus Topic: Solid-Phase Peptide Synthesis
- Notes
- Quiz Yourself!
- (Advanced) References and Further Reading
1. What Are Peptide Bonds?
A “peptide bond” is an amide linkage (see Amides: Properties. Synthesis, and Nomenclature) that connects two amino acids, as in the “dipeptides” L-phenylalanyl-L-valine (below left) and L-leucyl-L-alanine (below right):
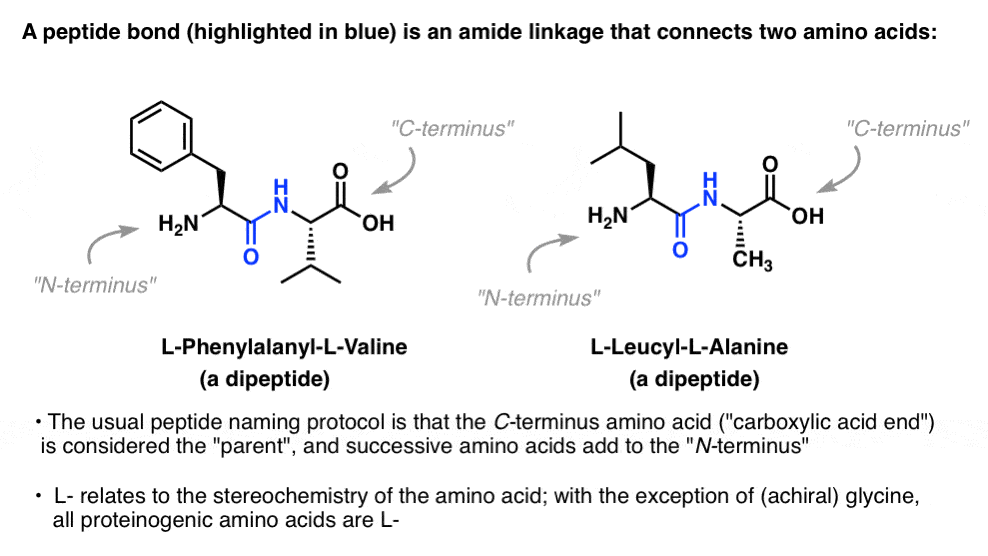
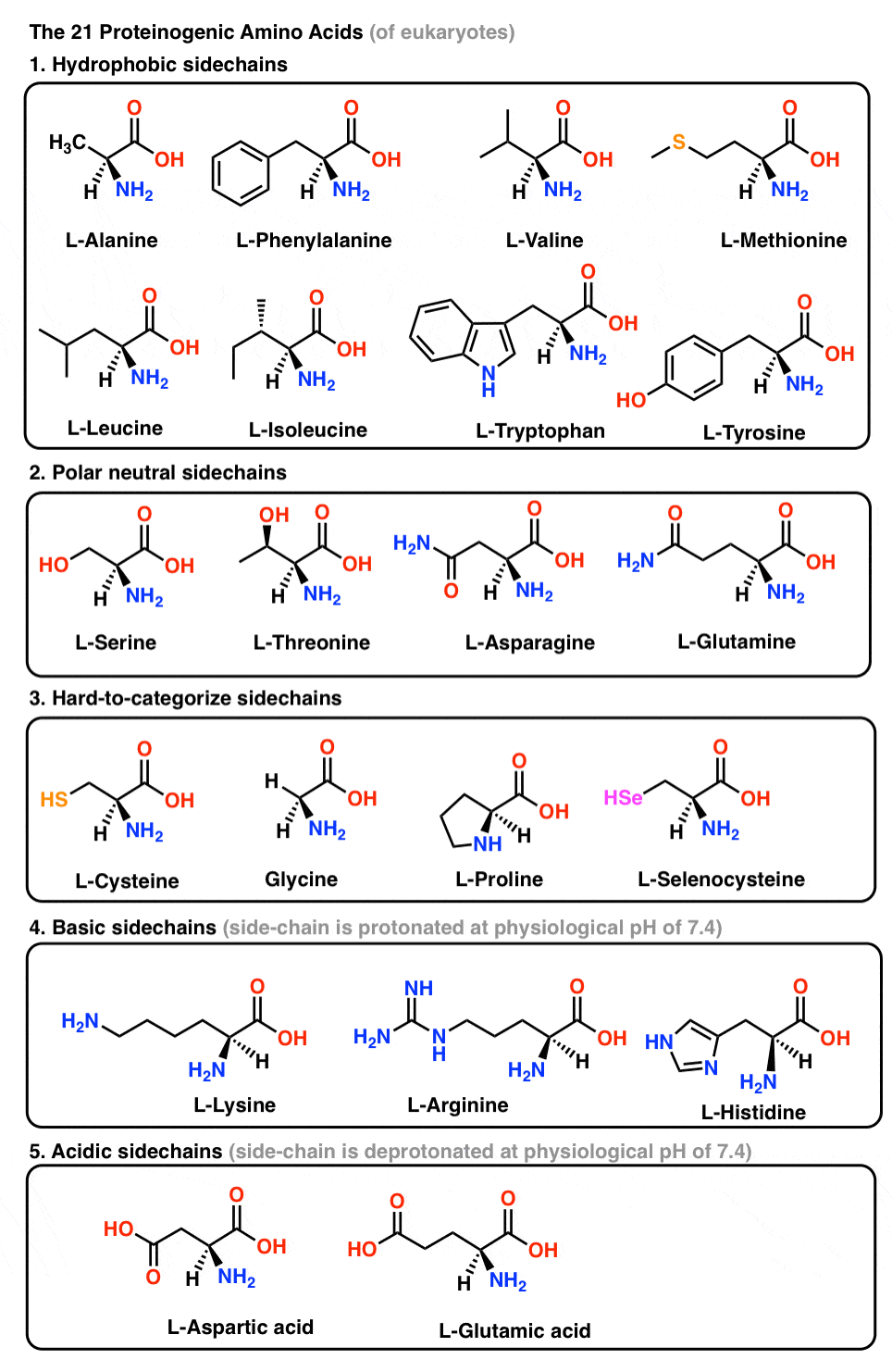
2. The “Proteinogenic” Amino Acids
Proteinogenic amino acids are the building blocks of proteins. In addition to the 20 amino acids directly encoded by the genome, two other amino acids are coded into proteins under special circumstances: selenocysteine (present in eukaryotes, including humans) and pyrrolysine (found only in methane-producing bacteria).
With the exception of (achiral) glycine, all proteinogenic amino acids are L-amino acids, where the “L-” prefix relates the stereochemistry of the amino acid relative to that of L-glyceraldehyde [See post: D and L Sugars] .
Of the chiral amino acids, all are S, with the exception of cysteine and selenocysteine [Note 1] (because sulfur and selenium have a higher priority under the CIP system. )
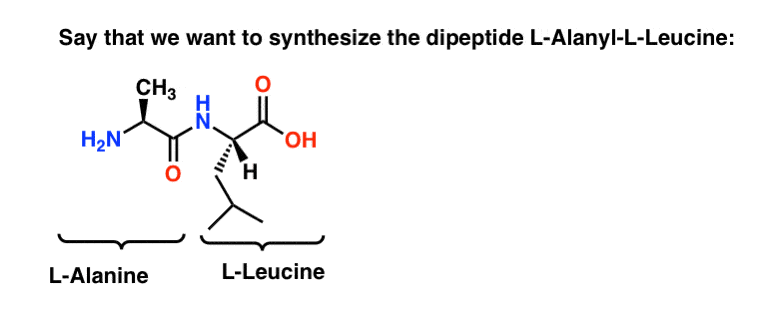
3. Synthesis of A Simple Dipeptide Without Protecting Groups (is not advisable!)
Let’s build a simple dipeptide between two of these amino acids. For simplicity’s sake, we’ll pick two from the “hydrophobic sidechain” group, alanine (Ala) and leucine (Leu), since their sidechains don’t need additional protecting groups.
What do we need to do to make L-Ala-L-Leu ?
Surveying the methods previously covered to make amides, it might seem simple.
Why not take 1 equivalent each of L-alanine and L-phenylalanine, add a coupling agent like N,N-dicyclohexylcarbodiimide (DCC) and patiently wait for our product to appear?
What could possibly go wrong?
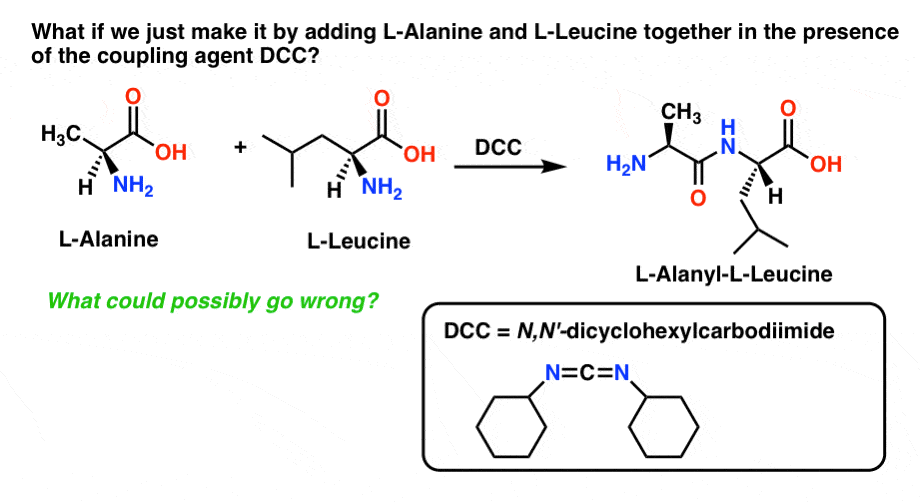
Well, this will give us some of our desired product. But it won’t do so efficiently!
That’s because each amino acid has two reactive termini – an amine and a carboxylic acid – and they can bond together in multiple ways.
Just like the letters A and L can combine to make the words AL and LA, in addition to Ala-Leu (our desired product) we will also get Leu-Ala.
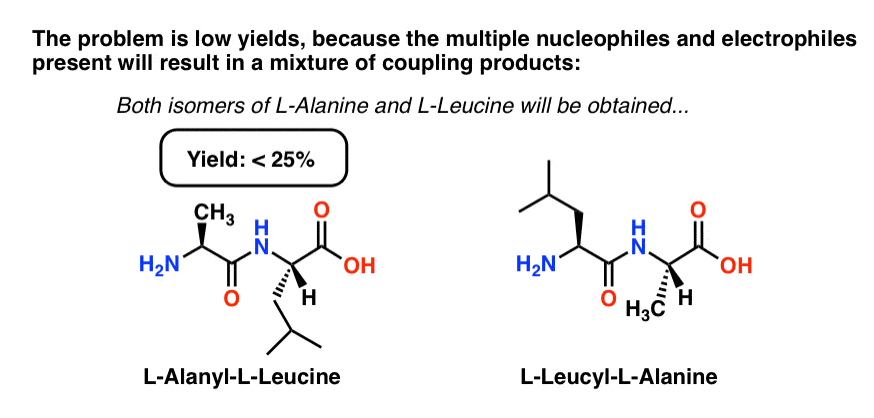
Furthermore, since we’re not adding single molecules together but molar quantities (even a millionth of a mole (a “micromole”) has 1017 molecules in it) we also have the possibility of forming the “homo-dipeptides” AA (Ala-Ala) and LL (Leu-Leu).
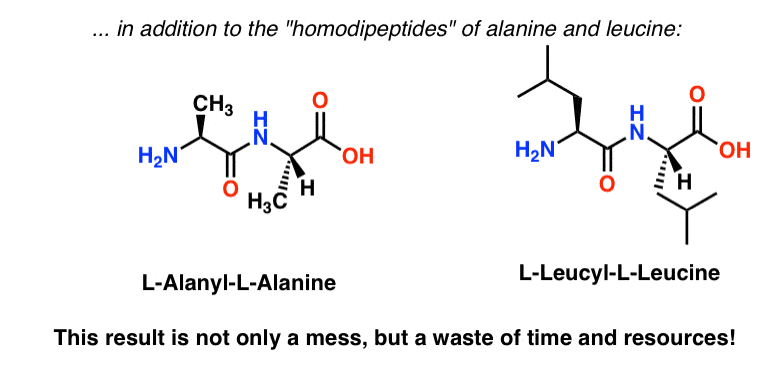
And that’s just a start. No matter how you slice it you’re looking at a low yield (<25%) of the desired material.
That’s inefficient, wasteful, and expensive! Isn’t there a better way?
4. Synthesis Of A Dipeptide Using A Protecting Group Strategy
Yes. Rather than using the native amino acids and just praying for a good yield, we can use protected versions of L-Alanine and L-Leucine.
If we protect the carboxylic acid of Leucine as an ester (e.g. a methyl ester) and protect the amine of L-Alanine as a carbamate (See: Carbamates as protecting groups) then we set up a situation where we have a single nucleophile and a single electrophile.
This results in a high yield (>95%) of a single product!
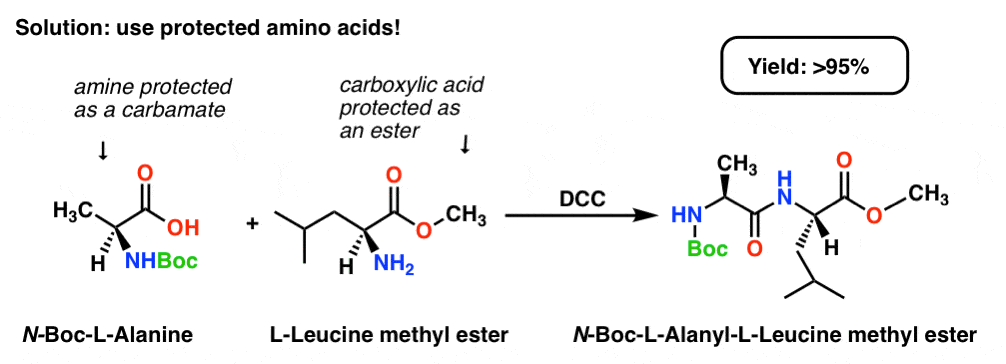
[Note: the Boc group is a popular carbamate protecting group for amines; “Boc” stands for t-butyloxycarbonyl]
5. Synthesis of Longer Peptides – Tripeptides and Tetrapeptides
The good news is that we don’t have to stop at the dipeptide. If we choose protecting groups that can be removed selectively (and the carbamate / ester pair qualifies) then we can then deprotect the carbamate, and add a third amino acid.
The choice of carbamate protecting group here was t-butoxycarbonyl (Boc) which is removed with strong acid (trifluoroacetic acid, abbreviated as TFA).
Treatment with TFA removes the Boc group but leaves the methyl ester alone.
So if we treat the dipeptide with TFA, we liberate the amine nitrogen, and can react with another Boc-protected amino acid in the presence of DCC to get a tripeptide.
If we’re keen, we can even extend the same method to build a tetrapeptide, a pentapeptide… or beyond!
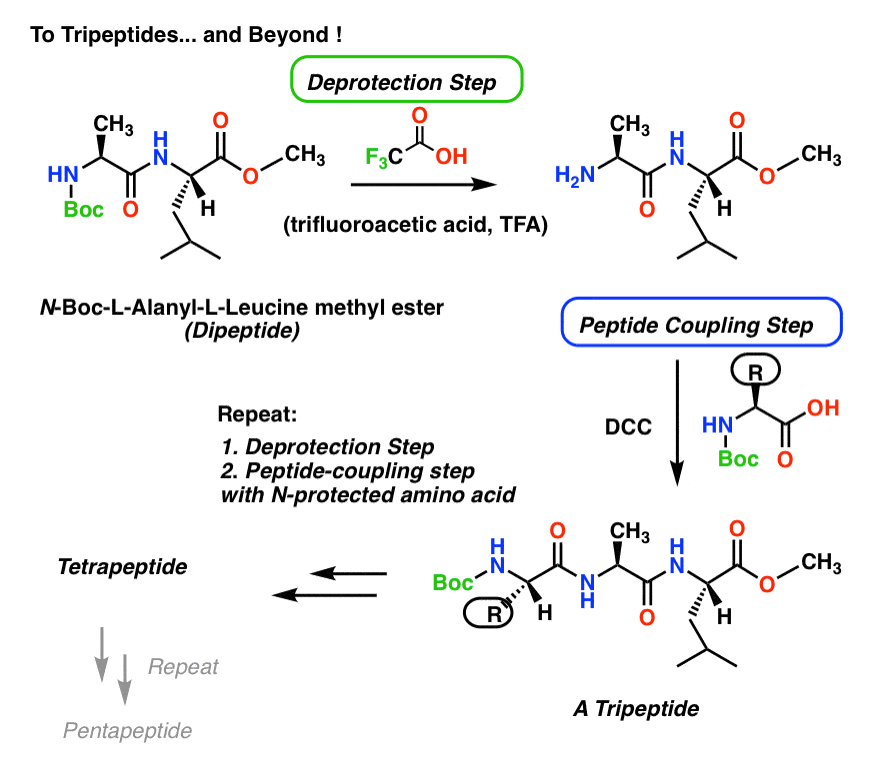
It’s not unreasonable to consider this method for longer peptides.
For instance, take something like bradykinin, a 9-peptide chain that causes dilation of blood vessels leading to a rapid drop in blood pressure. (Your body releases bradykinin in response to snake bites, which is how it was originally discovered.)
It might be interesting to synthesize variants of bradykinin where some of the amino acids are swapped out for other ones. In order to do that, we’d need to be able to synthesize it.
So how effective could it be?
If each peptide coupling step has a yield of about 95%, then our overall yield for making bradykinin would be (0.95)9 , or 63%. That’s actually pretty good! A lot of chemists would be happy to get a yield of 63% for a single reaction, let me tell you.
If the yields are high enough, one can even imagine building something crazy like insulin (51 peptide residues). That’s 7% yield for 51 steps.
Is this possible?
Yes… but it requires a clever modification that won its inventor, Bruce Merrifield, the 1984 Nobel Prize in Chemistry.
6. Bonus Topic: Solid-Phase Peptide Synthesis
What follows below is more supplemental than anything else, but given the importance of the topic, both interesting and useful.
In 1963 a chemist at Rockefeller University named Bruce Merrifield published a paper that would revolutionize how peptides were synthesized, and eventually make the synthesis of long peptides routine.
It was entitled: “Solid-Phase Peptide Synthesis. I. The Synthesis of a Tetrapeptide“.
Here’s the key idea.
Recall that in our original scheme (above) we protected the carboxylic acid as a methyl ester, which stays the same throughout the whole peptide synthesis.
Merrifield’s idea was: what if we find a way to attach the carboxylic acid to a functional group that is itself linked to a polymer bead? Not only would this also protect the carboxylic acid, it would drastically improve the ease of separations.
Why? Because instead of having to purify the final product by crystallization or column chromatography, you purify by filtering off the polymer beads (each 200-500 μm) and washing them to remove excess reagents.
The polymer beads themselves are pretty small. A typical size is 200 micrometers. Each bead can load about 4 nanomoles of amino acid.
[Brandon Finlay from ChemTips shows a picture here]
The video posted below is not mine, but it gives you an idea of the process.
At about 0:34 you can see how small the beads are.
The starting point for the Merrifield process is crosslinked polystyrene. which behaves like one big interlinked molecule. Polystyrene is then attached to a linker, which usually terminates with an NH2 group. This itself is usually protected; in order to activate the linker, you need to remove the protecting group cap.
The polymer bead needs to swell in a solvent in order for functional groups on the solid support to undergo reactions efficiently.
The essential procedure is: swell –> add reagents –> wait –> filter –> wash, and repeat. Beads stay in the reaction vessel the whole time. There’s also usually some kind of capping step to make sure any unreacted amines don’t participate in the next reaction.
It’s possible to make peptides up to about 50 units this way. In highly automated systems one can be even more ambitious.
Merrifield started knocking off peptides in the 1960s. Bradykinin was made in 8 days and 68% overall yield. one example. Insulin was made two years later. The crowning achievement of this initial period was probably ribonuclease A, which has 150 amino acid residues.
The original Merrifield process has been significantly modified and improved. Originally, removal of the linker required harsh conditions (strong acid). Today, procedures usually employ FMOC protecting groups instead of BOC, which allow for deprotection with mild amine base (piperidine). A galaxy of new resins, linkers, and coupling procedures have been subsequently developed. The Wikipedia article on solid-phase peptide synthesis is an OK place to start.
Notes
Related Articles
- Reactions of Diazonium Salts: Sandmeyer and Related Reactions
- Protecting Groups for Amines – Carbamates
- The Amide Functional Group: Properties, Synthesis, and Nomenclature
- Formation of Amides Using DCC (MOC Reaction Guide)
- Amine Protection and Deprotection (MOC Reaction Guide)
- Amine Practice Questions (MOC Membership)
- Isoelectric Points of Amino Acids (and How To Calculate Them)
Note 1. Cysteine (and selenocysteine) are L, but their stereocenters are (R), because sulfur and selenium have higher priority within the Cahn-Ingold-Prelog system.
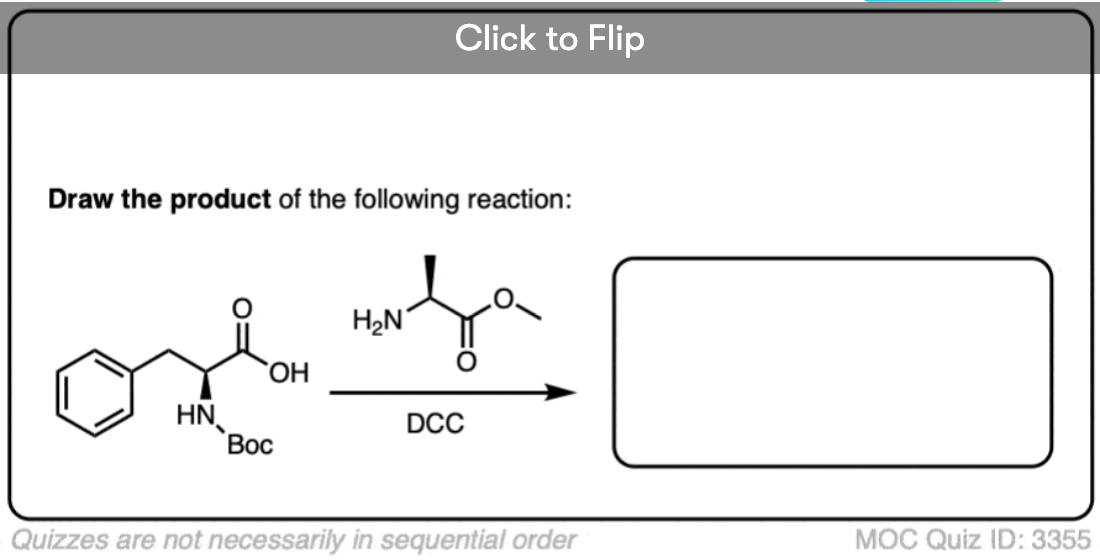
Quiz Yourself!
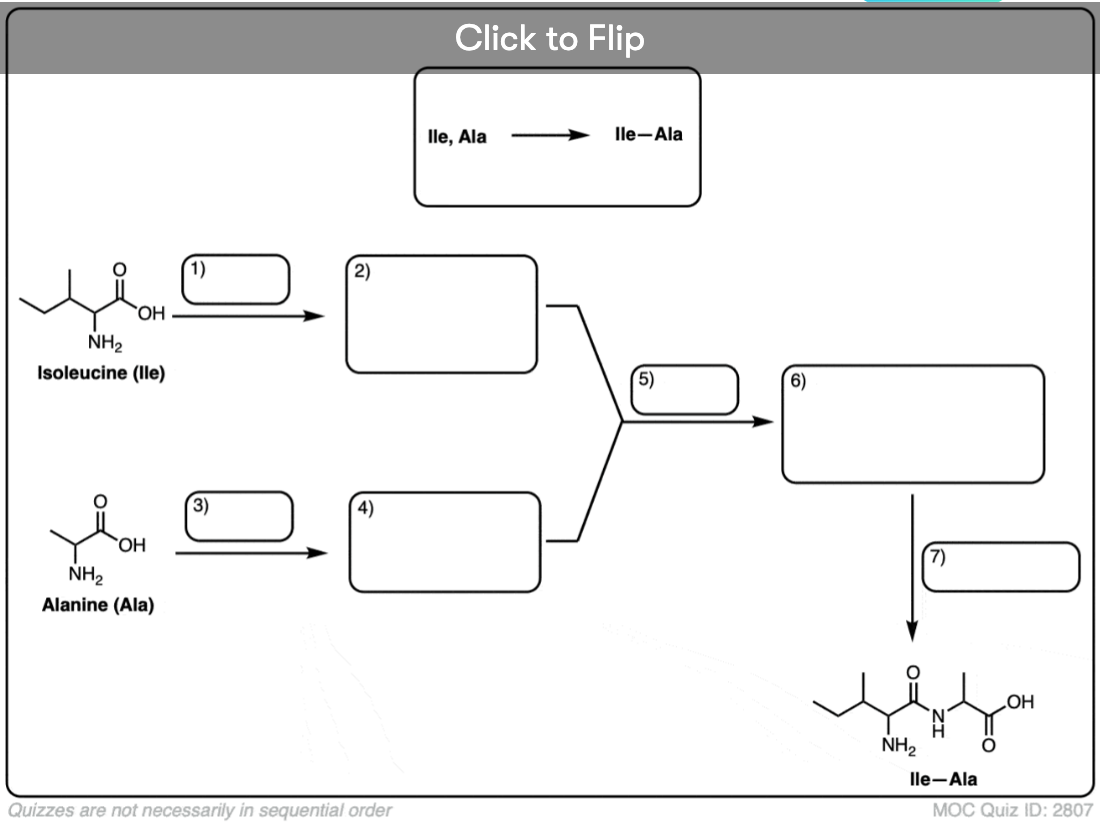
Become a MOC member to see the clickable quiz with answers on the back.
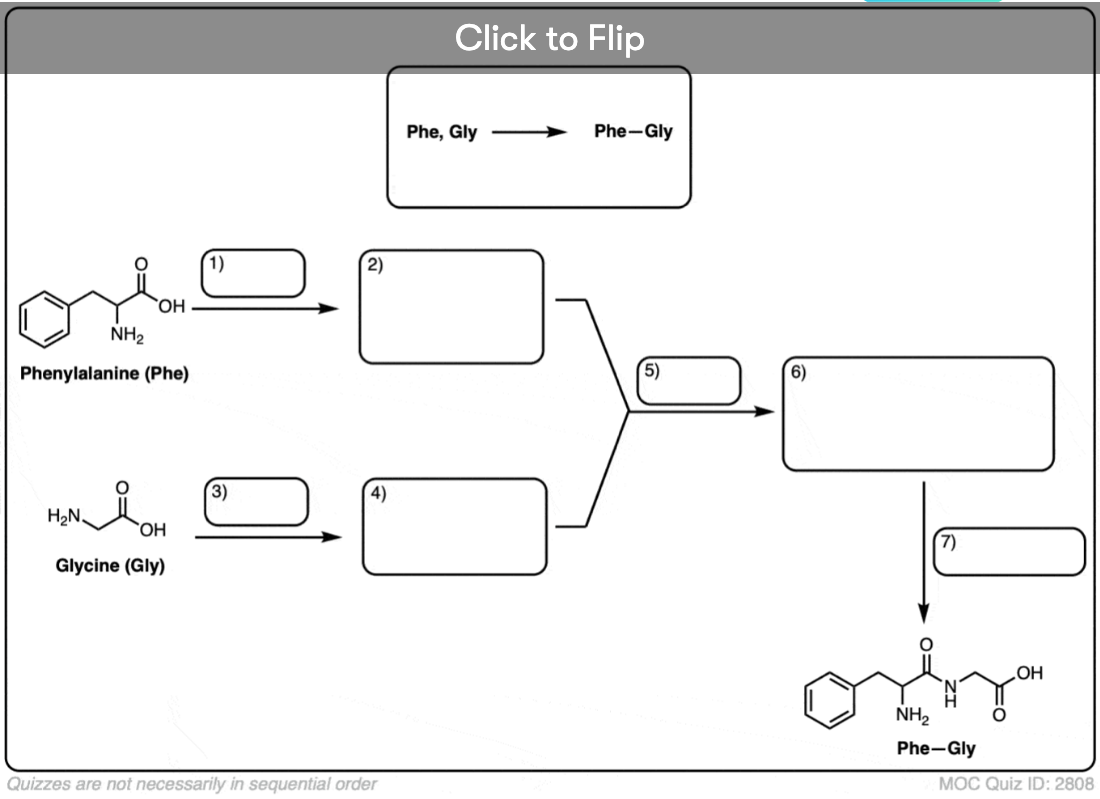
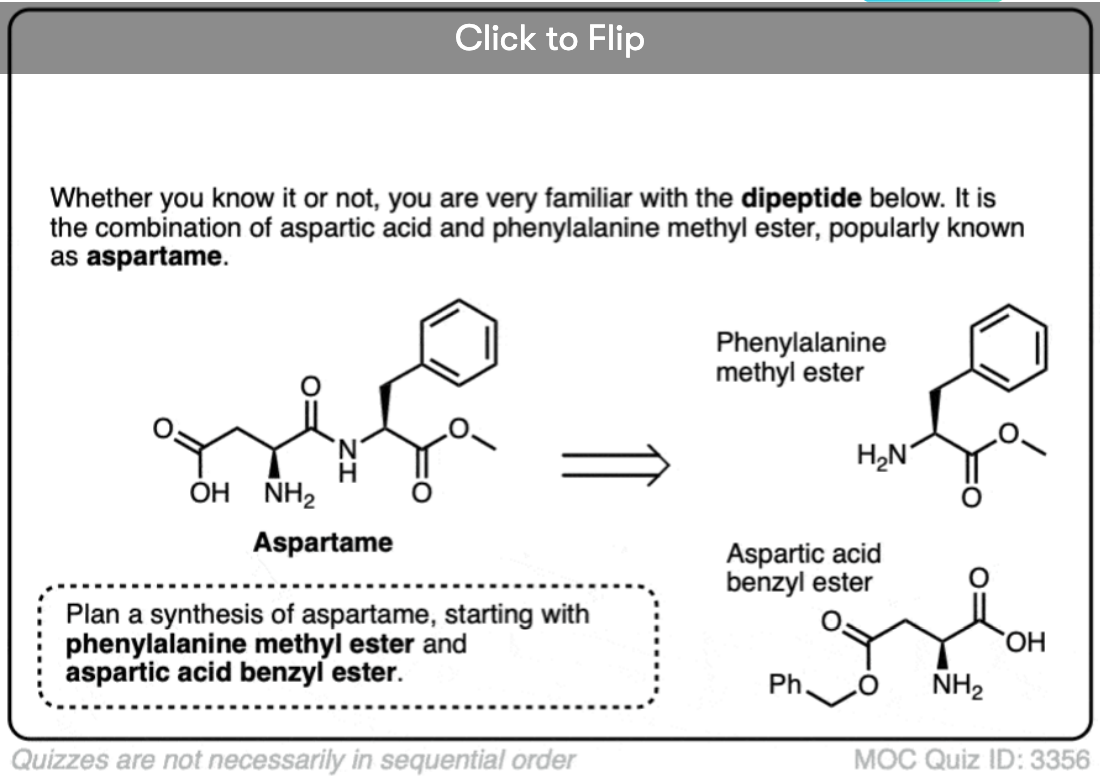
Become a MOC member to see the clickable quiz with answers on the back.
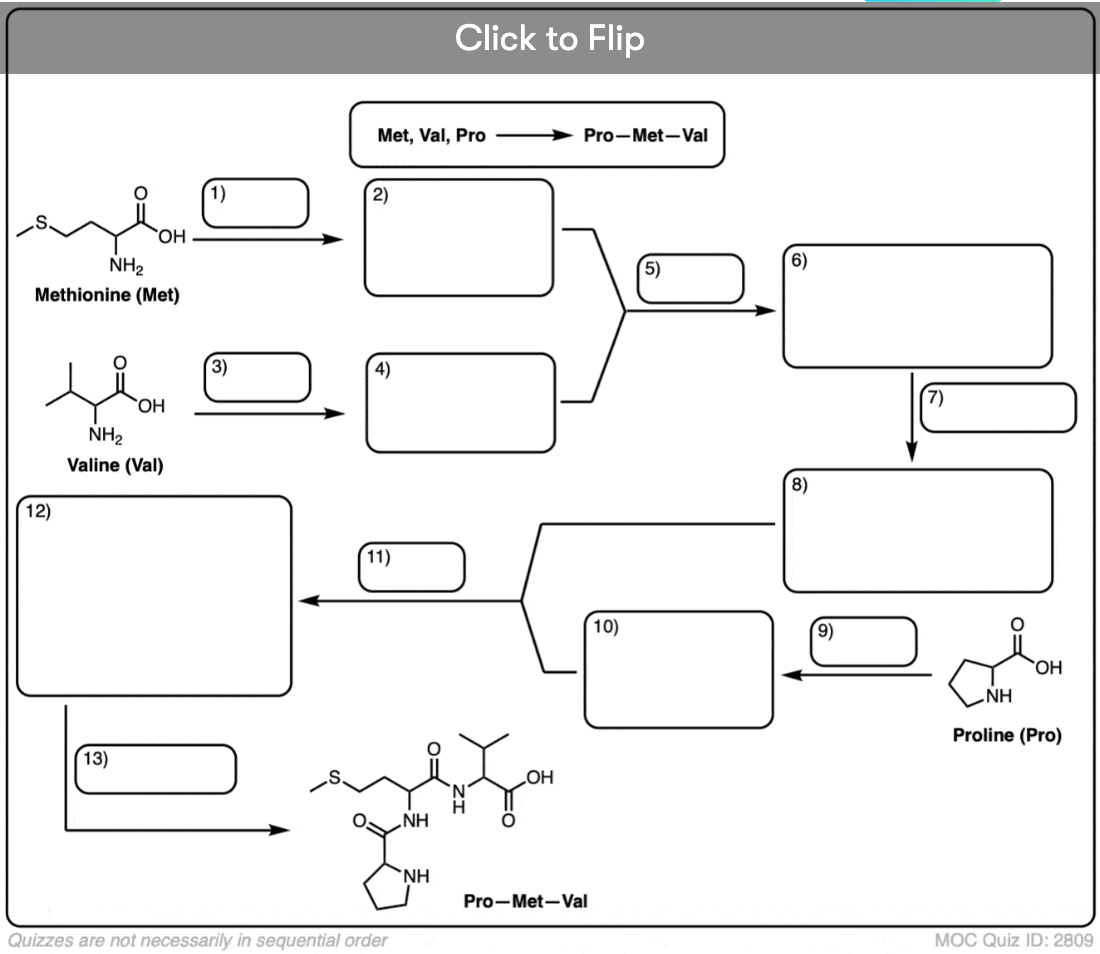
Become a MOC member to see the clickable quiz with answers on the back.
Become a MOC member to see the clickable quiz with answers on the back.
Become a MOC member to see the clickable quiz with answers on the back.
(Advanced) References and Further Reading
This is a major topic, as the synthesis of peptides is a global billion-dollar industry.
- THE SYNTHESIS OF AN OCTAPEPTIDE AMIDE WITH THE HORMONAL ACTIVITY OF OXYTOCIN
Vincent du Vigneaud, Charlotte Ressler, College John M. Swan, Carleton W. Roberts, Panayotis G. Katsoyannis, and Samuel Gordon
Journal of the American Chemical Society 1953, 75 (19), 4879-4880
DOI: 1021/ja01115a553 - The Synthesis of Oxytocin
Vincent du Vigneaud, Charlotte Ressler, John M. Swan, Carleton W. Roberts, and Panayotis G. Katsoyannis
Journal of the American Chemical Society 1954, 76 (12), 3115-3121
DOI: 1021/ja01641a004 - A Method of Synthesis of Long Peptide Chains Using a Synthesis of Oxytocin as an Example
Miklos Bodanszky and Vincent du Vigneaud
Journal of the American Chemical Society 1959, 81 (21), 5688-5691
DOI: 1021/ja01530a040
In the first half of the 20th century, peptide synthesis was done using standard organic chemistry solution phase techniques. This is now known as LPPS (liquid-phase peptide synthesis). du Vigneaud received the Nobel Prize in chemistry in 1955 for his work in showing that peptide synthesis could be achieved, using the correct choice of protecting groups and synthetic strategies.In 1963, Prof. Robert Bruce Merrifield (Rockefeller U., New York) revolutionized peptide synthesis by coming up with the SPPS (Solid-Phase Peptide Synthesis) method, making the synthesis of long peptide chains much more feasible. The C-terminus is bound to a polymer resin, and the amino acids are added one at a time following the same cycle: deprotect, wash, couple the next amino acid (with a peptide-coupling reagent such as DCC), wash, deprotect the N-terminus again, and so on. Merrifield’s method came to be called the Boc/Bzl strategy due to the protecting groups employed (Boc for the nitrogen atoms, and Bzl (benzyl) for the side chains). The catch is that final cleavage of the peptide from the resin using this method requires anhydrous HF, which is not easy to handle. - Solid Phase Peptide Synthesis. I. The Synthesis of a Tetrapeptide
R. B. Merrifield
Journal of the American Chemical Society 1963, 85 (14), 2149-2154
DOI: 10.1021/ja00897a025
This is the paper that started it all – a single-author publication by Prof. Merrifield using the SPPS method to make a tetrapeptide. This remains one of the most highly-viewed and highly-cited papers in JACS, even today. - The Synthesis of Bovine Insulin by the Solid Phase Method
Marglin and R. B. Merrifield
Journal of the American Chemical Society 1966, 88 (21), 5051-5052
DOI: 10.1021/ja00973a068
Insulin is a billion-dollar hormone as its dysregulation is what causes diabetes. This paper shows that insulin can be made through SPPS methods. Insulin is tricky to make as it has 2 chains (the A and B chain) connected through disulfide bonds. Interestingly, Merrifield synthesized the active hormone by combining both chains with protected thiols (protected as sulfonates), which he then reduced to the thiol, and then oxidized in air in a basic medium (pH 10). This is called undirected or air oxidation since the disulfide bonds are not being formed selectively; luckily they formed correctly in this case. Insulin today is not manufactured by the SPPS method due to these complications; instead it is made through a recombinant process. - The Synthesis of Ribonuclease A
Bernd Gutte and R. B. Merrifield
Journal of Biological Chemistry 1971, 246 (6), 1922-1941
DOI: 10.1016/S0021-9258(18)62396-8
This is the crowning achievement of SPPS – the synthesis of a 124-mer peptide (or protein, at this point), RNAse A. - The Solid Phase Synthesis of Ribonuclease A by Robert Bruce Merrifield
Nicole Kresge, Robert D. Simoni and Robert L. Hill
Journal of Biological Chemistry 2006, 281 (26), e21
DOI: 10.1016/S0021-9258(20)55702-5
A short biographical account of Prof. Merrifield’s life. This mentions that he developed the first prototype of an automated peptide synthesizer working in the basement of his house in 1965! - Solid Phase Synthesis
Nobel Lecture, December 8, 1984 by Bruce Merrifield
Merrifield’s Nobel Lecture upon receiving the Nobel Prize in Chemistry in 1984. This describes his life, how he conceived of and developed the SPPS process, and all the breakthroughs it has enabled. - 9-Fluorenylmethoxycarbonyl function, a new base-sensitive amino-protecting group
Louis A. Carpino and Grace Y. Han
Journal of the American Chemical Society 1970, 92 (19), 5748-5749
DOI: 10.1021/ja00722a043 - 9-Fluorenylmethoxycarbonyl amino-protecting group
Louis A. Carpino and Grace Y. Han
The Journal of Organic Chemistry 1972 37 (22), 3404-3409
DOI: 10.1021/jo00795a005
The discovery and development of the Fmoc (9-fluorenylmethoxycarbonyl) group as a protecting group for amines has also improved the practice of peptide synthesis and SPPS. - A mild procedure for solid phase peptide synthesis: use of fluorenylmethoxycarbonylamino-acids
Atherton, Hazel Fox, Diana Harkiss, C. J. Logan, R. C. Sheppard and B. J. Williams
J. Chem. Soc. Chem. Comm. 1978, 537-539
DOI: 10.1039/C39780000537
This is the first paper describing what is now known as the Fmoc/tBu SPPS process, which has largely supplanted the original Boc/Bzl process developed by Prof. Merrifield. The advantages with Fmoc-SPPS are multifold, but the main ones are simplicity and orthogonality. Amine deprotection is done with a base (20% piperidine in DMF is sufficient to deprotect an Fmoc-amine), and final cleavage of the peptide from the resin can be done with TFA (trifluoroacetic acid), which is much easier to handle than HF. - Advances in Fmoc solid-phase peptide synthesis
Raymond Behrendt, Peter White, and John Offer
Peptide Sci. 2016, 22, 4-27
DOI: 10.1002/psc.2836
A modern review on Fmoc-SPPS that describes how far we have come and some of the challenges that remain. For instance, aggregation of peptide chains on the resin is a major issue in Fmoc-SPPS when synthesizing especially hydrophobic peptides, and there are ways to deal with it, such as the introduction of ‘kinking’ residues, like Pro or the use of pseudoprolines, which will revert back to the desired amino acids when the peptide is cleaved with TFA. Native Chemical Ligation is also used nowadays for the synthesis of especially long peptides, like Merrifield’s RNAse A.
00 General Chemistry Review
01 Bonding, Structure, and Resonance
- How Do We Know Methane (CH4) Is Tetrahedral?
- Hybrid Orbitals and Hybridization
- How To Determine Hybridization: A Shortcut
- Orbital Hybridization And Bond Strengths
- Sigma bonds come in six varieties: Pi bonds come in one
- Dipole Moments and Dipoles
- A Key Skill: How to Calculate Formal Charge
- The Four Intermolecular Forces and How They Affect Boiling Points
- 3 Trends That Affect Boiling Points
- How To Use Electronegativity To Determine Electron Density (and why NOT to trust formal charge)
- Introduction to Resonance
- How To Use Curved Arrows To Interconvert Resonance Structures
- Evaluating Resonance Structures - The Fewer Charges, The Better
- How To Find The Best Resonance Structure By Applying Electronegativity
- Resonance Structures Should Have Full Octets (If Possible!)
- Resonance Structures: Negative Charges Are Best Placed On The Least Basic Atom
- Pi-Donation and Pi-Donors
- Exploring Resonance: Pi-acceptors
- In Summary: How To Evaluate Resonance Structures
- Drawing Resonance Structures: 3 Common Mistakes To Avoid
- How to apply electronegativity and resonance to understand reactivity
- Bond Hybridization Practice
- Structure and Bonding Practice Quizzes
- Resonance Structures Practice
02 Acid Base Reactions
- Introduction to Acid-Base Reactions
- Acid Base Reactions In Organic Chemistry
- Five Key Factors That Influence Acidity
- Factors That Affect Base Strength In Organic Chemistry
- Acid-Base Reactions: Introducing Ka and pKa
- How to Use a pKa Table
- The pKa Table Is Your Friend
- A Handy Rule of Thumb for Acid-Base Reactions
- Acid Base Reactions Are Fast
- pKa Values Span 60 Orders Of Magnitude
- How Protonation and Deprotonation Affect Reactivity
- Acid Base Practice Problems
03 Alkanes and Nomenclature
- Meet the (Most Important) Functional Groups
- Condensed Formulas: Deciphering What the Brackets Mean
- Hidden Hydrogens, Hidden Lone Pairs, Hidden Counterions
- Don't Be Futyl, Learn The Butyls
- Primary, Secondary, Tertiary, Quaternary In Organic Chemistry
- Branching, and Its Affect On Melting and Boiling Points
- The Many, Many Ways of Drawing Butane
- Wedge And Dash Convention For Tetrahedral Carbon
- Common Mistakes in Organic Chemistry: Pentavalent Carbon
- Table of Functional Group Priorities for Nomenclature
- Summary Sheet - Alkane Nomenclature
- Organic Chemistry IUPAC Nomenclature Demystified With A Simple Puzzle Piece Approach
- Boiling Point Quizzes
- Organic Chemistry Nomenclature Quizzes
04 Conformations and Cycloalkanes
- Staggered vs Eclipsed Conformations of Ethane
- Conformational Isomers of Propane
- Newman Projection of Butane (and Gauche Conformation)
- Introduction to Cycloalkanes
- Geometric Isomers In Small Rings: Cis And Trans Cycloalkanes
- Calculation of Ring Strain In Cycloalkanes
- Cycloalkanes - Ring Strain In Cyclopropane And Cyclobutane
- Cyclohexane Conformations
- Cyclohexane Chair Conformation: An Aerial Tour
- How To Draw The Cyclohexane Chair Conformation
- The Cyclohexane Chair Flip
- The Cyclohexane Chair Flip - Energy Diagram
- Substituted Cyclohexanes - Axial vs Equatorial
- Ranking The Bulkiness Of Substituents On Cyclohexanes: "A-Values"
- Cyclohexane Chair Conformation Stability: Which One Is Lower Energy?
- Fused Rings - Cis-Decalin and Trans-Decalin
- Naming Bicyclic Compounds - Fused, Bridged, and Spiro
- Bredt's Rule (And Summary of Cycloalkanes)
- Newman Projection Practice
- Cycloalkanes Practice Problems
05 A Primer On Organic Reactions
- The Most Important Question To Ask When Learning a New Reaction
- Curved Arrows (for reactions)
- Nucleophiles and Electrophiles
- The Three Classes of Nucleophiles
- Nucleophilicity vs. Basicity
- What Makes A Good Nucleophile?
- What Makes A Good Leaving Group?
- 3 Factors That Stabilize Carbocations
- Equilibrium and Energy Relationships
- 7 Factors that stabilize negative charge in organic chemistry
- 7 Factors That Stabilize Positive Charge in Organic Chemistry
- What's a Transition State?
- Hammond's Postulate
- Learning Organic Chemistry Reactions: A Checklist (PDF)
06 Free Radical Reactions
- Free Radical Reactions
- 3 Factors That Stabilize Free Radicals
- Bond Strengths And Radical Stability
- Free Radical Initiation: Why Is "Light" Or "Heat" Required?
- Initiation, Propagation, Termination
- Monochlorination Products Of Propane, Pentane, And Other Alkanes
- Selectivity In Free Radical Reactions
- Selectivity in Free Radical Reactions: Bromination vs. Chlorination
- Halogenation At Tiffany's
- Allylic Bromination
- Bonus Topic: Allylic Rearrangements
- In Summary: Free Radicals
- Synthesis (2) - Reactions of Alkanes
- Free Radicals Practice Quizzes
07 Stereochemistry and Chirality
- Types of Isomers: Constitutional Isomers, Stereoisomers, Enantiomers, and Diastereomers
- How To Draw The Enantiomer Of A Chiral Molecule
- How To Draw A Bond Rotation
- Introduction to Assigning (R) and (S): The Cahn-Ingold-Prelog Rules
- Assigning Cahn-Ingold-Prelog (CIP) Priorities (2) - The Method of Dots
- Enantiomers vs Diastereomers vs The Same? Two Methods For Solving Problems
- Assigning R/S To Newman Projections (And Converting Newman To Line Diagrams)
- How To Determine R and S Configurations On A Fischer Projection
- The Meso Trap
- Optical Rotation, Optical Activity, and Specific Rotation
- Optical Purity and Enantiomeric Excess
- What's a Racemic Mixture?
- Chiral Allenes And Chiral Axes
- Stereochemistry Practice Problems and Quizzes
08 Substitution Reactions
- Nucleophilic Substitution Reactions - Introduction
- Two Types of Nucleophilic Substitution Reactions
- The SN2 Mechanism
- Why the SN2 Reaction Is Powerful
- The SN1 Mechanism
- The Conjugate Acid Is A Better Leaving Group
- Comparing the SN1 and SN2 Reactions
- Polar Protic? Polar Aprotic? Nonpolar? All About Solvents
- Steric Hindrance is Like a Fat Goalie
- Common Blind Spot: Intramolecular Reactions
- Substitution Practice - SN1
- Substitution Practice - SN2
09 Elimination Reactions
- Elimination Reactions (1): Introduction And The Key Pattern
- Elimination Reactions (2): The Zaitsev Rule
- Elimination Reactions Are Favored By Heat
- Two Elimination Reaction Patterns
- The E1 Reaction
- The E2 Mechanism
- E1 vs E2: Comparing the E1 and E2 Reactions
- Antiperiplanar Relationships: The E2 Reaction and Cyclohexane Rings
- Bulky Bases in Elimination Reactions
- Comparing the E1 vs SN1 Reactions
- Elimination (E1) Reactions With Rearrangements
- E1cB - Elimination (Unimolecular) Conjugate Base
- Elimination (E1) Practice Problems And Solutions
- Elimination (E2) Practice Problems and Solutions
10 Rearrangements
11 SN1/SN2/E1/E2 Decision
- Identifying Where Substitution and Elimination Reactions Happen
- Deciding SN1/SN2/E1/E2 (1) - The Substrate
- Deciding SN1/SN2/E1/E2 (2) - The Nucleophile/Base
- SN1 vs E1 and SN2 vs E2 : The Temperature
- Deciding SN1/SN2/E1/E2 - The Solvent
- Wrapup: The Key Factors For Determining SN1/SN2/E1/E2
- Alkyl Halide Reaction Map And Summary
- SN1 SN2 E1 E2 Practice Problems
12 Alkene Reactions
- E and Z Notation For Alkenes (+ Cis/Trans)
- Alkene Stability
- Alkene Addition Reactions: "Regioselectivity" and "Stereoselectivity" (Syn/Anti)
- Stereoselective and Stereospecific Reactions
- Hydrohalogenation of Alkenes and Markovnikov's Rule
- Hydration of Alkenes With Aqueous Acid
- Rearrangements in Alkene Addition Reactions
- Halogenation of Alkenes and Halohydrin Formation
- Oxymercuration Demercuration of Alkenes
- Hydroboration Oxidation of Alkenes
- m-CPBA (meta-chloroperoxybenzoic acid)
- OsO4 (Osmium Tetroxide) for Dihydroxylation of Alkenes
- Palladium on Carbon (Pd/C) for Catalytic Hydrogenation of Alkenes
- Cyclopropanation of Alkenes
- A Fourth Alkene Addition Pattern - Free Radical Addition
- Alkene Reactions: Ozonolysis
- Oxidative Cleavage of Vicinal Diols With NaIO4 and Pb(OAc)4
- Summary: Three Key Families Of Alkene Reaction Mechanisms
- Synthesis (4) - Alkene Reaction Map, Including Alkyl Halide Reactions
- Alkene Reactions Practice Problems
13 Alkyne Reactions
- Acetylides from Alkynes, And Substitution Reactions of Acetylides
- Partial Reduction of Alkynes With Lindlar's Catalyst
- Partial Reduction of Alkynes With Na/NH3 To Obtain Trans Alkenes
- Alkyne Hydroboration With "R2BH"
- Hydration and Oxymercuration of Alkynes
- Hydrohalogenation of Alkynes
- Alkyne Halogenation: Bromination and Chlorination of Alkynes
- Oxidation of Alkynes With O3 and KMnO4
- Alkenes To Alkynes Via Halogenation And Elimination Reactions
- Alkynes Are A Blank Canvas
- Synthesis (5) - Reactions of Alkynes
- Alkyne Reactions Practice Problems With Answers
14 Alcohols, Epoxides and Ethers
- Alcohols - Nomenclature and Properties
- Alcohols Can Act As Acids Or Bases (And Why It Matters)
- Alcohols - Acidity and Basicity
- The Williamson Ether Synthesis
- Ethers From Alkenes, Tertiary Alkyl Halides and Alkoxymercuration
- Alcohols To Ethers via Acid Catalysis
- Cleavage Of Ethers With Acid
- Epoxides - The Outlier Of The Ether Family
- Opening of Epoxides With Acid
- Epoxide Ring Opening With Base
- Making Alkyl Halides From Alcohols
- Tosylates And Mesylates
- PBr3 and SOCl2
- Elimination Reactions of Alcohols
- Elimination of Alcohols To Alkenes With POCl3
- Alcohol Oxidation: "Strong" and "Weak" Oxidants
- Demystifying The Mechanisms of Alcohol Oxidations
- Protecting Groups For Alcohols
- Thiols And Thioethers
- Calculating the oxidation state of a carbon
- Oxidation and Reduction in Organic Chemistry
- Oxidation Ladders
- SOCl2 Mechanism For Alcohols To Alkyl Halides: SN2 versus SNi
- Alcohol Reactions Roadmap (PDF)
- Alcohol Reaction Practice Problems
- Epoxide Reaction Quizzes
- Oxidation and Reduction Practice Quizzes
15 Organometallics
- What's An Organometallic?
- Formation of Grignard and Organolithium Reagents
- Organometallics Are Strong Bases
- Reactions of Grignard Reagents
- Protecting Groups In Grignard Reactions
- Synthesis Problems Involving Grignard Reagents
- Grignard Reactions And Synthesis (2)
- Organocuprates (Gilman Reagents): How They're Made
- Gilman Reagents (Organocuprates): What They're Used For
- The Heck, Suzuki, and Olefin Metathesis Reactions (And Why They Don't Belong In Most Introductory Organic Chemistry Courses)
- Reaction Map: Reactions of Organometallics
- Grignard Practice Problems
16 Spectroscopy
- Degrees of Unsaturation (or IHD, Index of Hydrogen Deficiency)
- Conjugation And Color (+ How Bleach Works)
- Introduction To UV-Vis Spectroscopy
- UV-Vis Spectroscopy: Absorbance of Carbonyls
- UV-Vis Spectroscopy: Practice Questions
- Bond Vibrations, Infrared Spectroscopy, and the "Ball and Spring" Model
- Infrared (IR) Spectroscopy: A Quick Primer On Interpreting Spectra
- IR Spectroscopy: 4 Practice Problems
- 1H NMR: How Many Signals?
- Homotopic, Enantiotopic, Diastereotopic
- Diastereotopic Protons in 1H NMR Spectroscopy: Examples
- 13-C NMR - How Many Signals
- Liquid Gold: Pheromones In Doe Urine
- Natural Product Isolation (1) - Extraction
- Natural Product Isolation (2) - Purification Techniques, An Overview
- Structure Determination Case Study: Deer Tarsal Gland Pheromone
17 Dienes and MO Theory
- What To Expect In Organic Chemistry 2
- Are these molecules conjugated?
- Conjugation And Resonance In Organic Chemistry
- Bonding And Antibonding Pi Orbitals
- Molecular Orbitals of The Allyl Cation, Allyl Radical, and Allyl Anion
- Pi Molecular Orbitals of Butadiene
- Reactions of Dienes: 1,2 and 1,4 Addition
- Thermodynamic and Kinetic Products
- More On 1,2 and 1,4 Additions To Dienes
- s-cis and s-trans
- The Diels-Alder Reaction
- Cyclic Dienes and Dienophiles in the Diels-Alder Reaction
- Stereochemistry of the Diels-Alder Reaction
- Exo vs Endo Products In The Diels Alder: How To Tell Them Apart
- HOMO and LUMO In the Diels Alder Reaction
- Why Are Endo vs Exo Products Favored in the Diels-Alder Reaction?
- Diels-Alder Reaction: Kinetic and Thermodynamic Control
- The Retro Diels-Alder Reaction
- The Intramolecular Diels Alder Reaction
- Regiochemistry In The Diels-Alder Reaction
- The Cope and Claisen Rearrangements
- Electrocyclic Reactions
- Electrocyclic Ring Opening And Closure (2) - Six (or Eight) Pi Electrons
- Diels Alder Practice Problems
- Molecular Orbital Theory Practice
18 Aromaticity
- Introduction To Aromaticity
- Rules For Aromaticity
- Huckel's Rule: What Does 4n+2 Mean?
- Aromatic, Non-Aromatic, or Antiaromatic? Some Practice Problems
- Antiaromatic Compounds and Antiaromaticity
- The Pi Molecular Orbitals of Benzene
- The Pi Molecular Orbitals of Cyclobutadiene
- Frost Circles
- Aromaticity Practice Quizzes
19 Reactions of Aromatic Molecules
- Electrophilic Aromatic Substitution: Introduction
- Activating and Deactivating Groups In Electrophilic Aromatic Substitution
- Electrophilic Aromatic Substitution - The Mechanism
- Ortho-, Para- and Meta- Directors in Electrophilic Aromatic Substitution
- Understanding Ortho, Para, and Meta Directors
- Why are halogens ortho- para- directors?
- Disubstituted Benzenes: The Strongest Electron-Donor "Wins"
- Electrophilic Aromatic Substitutions (1) - Halogenation of Benzene
- Electrophilic Aromatic Substitutions (2) - Nitration and Sulfonation
- EAS Reactions (3) - Friedel-Crafts Acylation and Friedel-Crafts Alkylation
- Intramolecular Friedel-Crafts Reactions
- Nucleophilic Aromatic Substitution (NAS)
- Nucleophilic Aromatic Substitution (2) - The Benzyne Mechanism
- Reactions on the "Benzylic" Carbon: Bromination And Oxidation
- The Wolff-Kishner, Clemmensen, And Other Carbonyl Reductions
- More Reactions on the Aromatic Sidechain: Reduction of Nitro Groups and the Baeyer Villiger
- Aromatic Synthesis (1) - "Order Of Operations"
- Synthesis of Benzene Derivatives (2) - Polarity Reversal
- Aromatic Synthesis (3) - Sulfonyl Blocking Groups
- Birch Reduction
- Synthesis (7): Reaction Map of Benzene and Related Aromatic Compounds
- Aromatic Reactions and Synthesis Practice
- Electrophilic Aromatic Substitution Practice Problems
20 Aldehydes and Ketones
- What's The Alpha Carbon In Carbonyl Compounds?
- Nucleophilic Addition To Carbonyls
- Aldehydes and Ketones: 14 Reactions With The Same Mechanism
- Sodium Borohydride (NaBH4) Reduction of Aldehydes and Ketones
- Grignard Reagents For Addition To Aldehydes and Ketones
- Wittig Reaction
- Hydrates, Hemiacetals, and Acetals
- Imines - Properties, Formation, Reactions, and Mechanisms
- All About Enamines
- Breaking Down Carbonyl Reaction Mechanisms: Reactions of Anionic Nucleophiles (Part 2)
- Aldehydes Ketones Reaction Practice
21 Carboxylic Acid Derivatives
- Nucleophilic Acyl Substitution (With Negatively Charged Nucleophiles)
- Addition-Elimination Mechanisms With Neutral Nucleophiles (Including Acid Catalysis)
- Basic Hydrolysis of Esters - Saponification
- Transesterification
- Proton Transfer
- Fischer Esterification - Carboxylic Acid to Ester Under Acidic Conditions
- Lithium Aluminum Hydride (LiAlH4) For Reduction of Carboxylic Acid Derivatives
- LiAlH[Ot-Bu]3 For The Reduction of Acid Halides To Aldehydes
- Di-isobutyl Aluminum Hydride (DIBAL) For The Partial Reduction of Esters and Nitriles
- Amide Hydrolysis
- Thionyl Chloride (SOCl2) And Conversion of Carboxylic Acids to Acid Halides
- Diazomethane (CH2N2)
- Carbonyl Chemistry: Learn Six Mechanisms For the Price Of One
- Making Music With Mechanisms (PADPED)
- Carboxylic Acid Derivatives Practice Questions
22 Enols and Enolates
- Keto-Enol Tautomerism
- Enolates - Formation, Stability, and Simple Reactions
- Kinetic Versus Thermodynamic Enolates
- Aldol Addition and Condensation Reactions
- Reactions of Enols - Acid-Catalyzed Aldol, Halogenation, and Mannich Reactions
- Claisen Condensation and Dieckmann Condensation
- Decarboxylation
- The Malonic Ester and Acetoacetic Ester Synthesis
- The Michael Addition Reaction and Conjugate Addition
- The Robinson Annulation
- Haloform Reaction
- The Hell–Volhard–Zelinsky Reaction
- Enols and Enolates Practice Quizzes
23 Amines
- The Amide Functional Group: Properties, Synthesis, and Nomenclature
- Basicity of Amines And pKaH
- 5 Key Basicity Trends of Amines
- The Mesomeric Effect And Aromatic Amines
- Nucleophilicity of Amines
- Alkylation of Amines (Sucks!)
- Reductive Amination
- The Gabriel Synthesis
- Some Reactions of Azides
- The Hofmann Elimination
- The Hofmann and Curtius Rearrangements
- The Cope Elimination
- Protecting Groups for Amines - Carbamates
- The Strecker Synthesis of Amino Acids
- Introduction to Peptide Synthesis
- Reactions of Diazonium Salts: Sandmeyer and Related Reactions
- Amine Practice Questions
24 Carbohydrates
- D and L Notation For Sugars
- Pyranoses and Furanoses: Ring-Chain Tautomerism In Sugars
- What is Mutarotation?
- Reducing Sugars
- The Big Damn Post Of Carbohydrate-Related Chemistry Definitions
- The Haworth Projection
- Converting a Fischer Projection To A Haworth (And Vice Versa)
- Reactions of Sugars: Glycosylation and Protection
- The Ruff Degradation and Kiliani-Fischer Synthesis
- Isoelectric Points of Amino Acids (and How To Calculate Them)
- Carbohydrates Practice
- Amino Acid Quizzes
25 Fun and Miscellaneous
- A Gallery of Some Interesting Molecules From Nature
- Screw Organic Chemistry, I'm Just Going To Write About Cats
- On Cats, Part 1: Conformations and Configurations
- On Cats, Part 2: Cat Line Diagrams
- On Cats, Part 4: Enantiocats
- On Cats, Part 6: Stereocenters
- Organic Chemistry Is Shit
- The Organic Chemistry Behind "The Pill"
- Maybe they should call them, "Formal Wins" ?
- Why Do Organic Chemists Use Kilocalories?
- The Principle of Least Effort
- Organic Chemistry GIFS - Resonance Forms
- Reproducibility In Organic Chemistry
- What Holds The Nucleus Together?
- How Reactions Are Like Music
- Organic Chemistry and the New MCAT
26 Organic Chemistry Tips and Tricks
- Common Mistakes: Formal Charges Can Mislead
- Partial Charges Give Clues About Electron Flow
- Draw The Ugly Version First
- Organic Chemistry Study Tips: Learn the Trends
- The 8 Types of Arrows In Organic Chemistry, Explained
- Top 10 Skills To Master Before An Organic Chemistry 2 Final
- Common Mistakes with Carbonyls: Carboxylic Acids... Are Acids!
- Planning Organic Synthesis With "Reaction Maps"
- Alkene Addition Pattern #1: The "Carbocation Pathway"
- Alkene Addition Pattern #2: The "Three-Membered Ring" Pathway
- Alkene Addition Pattern #3: The "Concerted" Pathway
- Number Your Carbons!
- The 4 Major Classes of Reactions in Org 1
- How (and why) electrons flow
- Grossman's Rule
- Three Exam Tips
- A 3-Step Method For Thinking Through Synthesis Problems
- Putting It Together
- Putting Diels-Alder Products in Perspective
- The Ups and Downs of Cyclohexanes
- The Most Annoying Exceptions in Org 1 (Part 1)
- The Most Annoying Exceptions in Org 1 (Part 2)
- The Marriage May Be Bad, But the Divorce Still Costs Money
- 9 Nomenclature Conventions To Know
- Nucleophile attacks Electrophile
27 Case Studies of Successful O-Chem Students
- Success Stories: How Corina Got The The "Hard" Professor - And Got An A+ Anyway
- How Helena Aced Organic Chemistry
- From a "Drop" To B+ in Org 2 – How A Hard Working Student Turned It Around
- How Serge Aced Organic Chemistry
- Success Stories: How Zach Aced Organic Chemistry 1
- Success Stories: How Kari Went From C– to B+
- How Esther Bounced Back From a "C" To Get A's In Organic Chemistry 1 And 2
- How Tyrell Got The Highest Grade In Her Organic Chemistry Course
- This Is Why Students Use Flashcards
- Success Stories: How Stu Aced Organic Chemistry
- How John Pulled Up His Organic Chemistry Exam Grades
- Success Stories: How Nathan Aced Organic Chemistry (Without It Taking Over His Life)
- How Chris Aced Org 1 and Org 2
- Interview: How Jay Got an A+ In Organic Chemistry
- How to Do Well in Organic Chemistry: One Student's Advice
- "America's Top TA" Shares His Secrets For Teaching O-Chem
- "Organic Chemistry Is Like..." - A Few Metaphors
- How To Do Well In Organic Chemistry: Advice From A Tutor
- Guest post: "I went from being afraid of tests to actually looking forward to them".
Thanks for very good explanation and excellent step-by-step video!!
These tips are awesome, I was looking for something like that. Thanks!
OK. Do you guys actually do peptide synthesis or do you just drop ship?
We, or least, in my school do cover this topic extensively. We need to learn fischer esterfication of them, acylation, protecting group although it was different than the protecting group you used It had two benzene rings and a five member ring in the middle.
Good to know! Sounds like you’re talking about FMOC. https://en.wikipedia.org/wiki/Fluorenylmethyloxycarbonyl_protecting_group
I should update to include that.
Thank you very much!